Algorithms#
Methods Implemented#
Optking performs Newton-Raphson (NR) and Qausi-NR optimizations for small to medium-sized molecules.
Algorithms include:
- Minimization
Steepest descent
NR
Rational function optimization (RFO)
Restricted Step RFO
Conjugate Gradient
- TS
Image RFO
Partioned RFO
- Reaction Path
Intrinsic Reaction Coordinate (IRC)
The type of optimization is controlled by the step_type and opt_type keywords. step_type chooses optimization algorithm (SD, NR, etc.)
opt_type selects the kind of optimization (min, TS, or IRC) and unless overriden chooses the appropriate (or default) step_type.
Classes and Functions#
For more information on the OptHelper classes see OptHelper To simply run an optimization
the OptHelper classes or the interfaces through QCEngine and Psi4 are recommended.
For more information on interacting directly with the algorithms see the API documentation below. In short,
the OptimizationAlgorithm class provides a basic “interface” for running optimizations through the take_step()
method. The OptimizationManager class extends this interface to encompass the addition of linesearching
to any of the basic algorithms and IRCFollowing.
optking.stepAlgorithms Module#
Defines classes for minimization and transition state optimizations
Classes#
- OptimizationInterface
An abstract class that all optimization classes inherit from. Requires that all subclasses implement a
take_step()method- OptimizationAlgorithm
An abstract class that extends
OptimizationInterfacemost algorithms (minimization or ts) inherit from this class.- QuasiNewtonOptimization
Abstract class for performing Quasi-Newton optimizations, these methods utilize an approximate updated hessian to approximate the potential energy surface.
- RFO
An abstract class for performing various Rational Function Optimization. RFO itself is an extension of a QuasiNewtonOptimization.
- NewtonRaphson
Extension of QuasiNewtonOptimization
- RestrictedStepRFO
Default minimization algorithm.
- ImageRFO
Extension of RestrictedStepRFO for transition state finding
- PartitionedRFO
Default TS algorithm before OptKing
v0.3.0- SteepestDescent
Basic gradient descent
- ConjugateGradient
Three varieties of CG are implemented Fletcher, Descent, and Polak. Fletcher is default.
- Linesearch
1-D linesearch by quadratic fit of energies
Functions#
- step_matches_forces
A basic attempt to determine whether the step taken matches the symmetry of the gradient.
Notes
The optimization classes above may be created using the factory pattern through
optking.optimize.optimization_factory(). If linesearching or more advanced management of the optimization
process is desired an OptimizationManager should be created.
(More features are coming to the OptimizationManager)
Functions#
|
Check for eigenvectors from RFO matrix which don't qualitatively match the gradient. |
Classes#
|
Implements the conjugate gradient algorithm. |
|
|
|
The standard minimization and transition state algorithms inherit from here. |
|
Declares that ALL OptKing optimization methods/algorithms will have a self.take_step() method. |
|
Partitions the gradient augmented Hessian into eigenvectors to maximize along (direction of the TS) and minimize along all other directions. |
|
|
|
|
|
Standard RFO and base class for RS_RFO, P_RFO, RS_PRFO #TODO |
|
Rational Function approximation (or 2x2 Pade approximation) for step. |
|
Steepest descent with step size adjustment |
Class Inheritance Diagram#
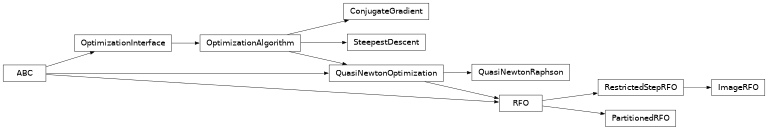
optking.linesearch Module#
Classes#
|
Basic framework for performing Linesearches. |
|
Extension of history.Step |
|
Class Inheritance Diagram#
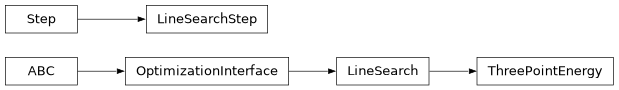
optking.IRCfollowing Module#
Functions#
|
Compute anticipated energy change along one dimension |
|
Computes distance scaling factor for mass-weighted internals. |
Classes#
|
Class Inheritance Diagram#